Doctor Yagiz Alagoz
Candidature
Graduated PhD 2020
Thesis Title
Feedback Regulation of Carotenoid Biosynthesis by an Apocarotenoid Sensing RNA Structural Switch
Research project
 Carotenoids are natural pigments synthesized by plants, algae, photosynthetic bacteria and aphids. In plants, carotenoids take role in photosynthesis, photoprotection and production of some phytohormones (Abscisic acid and Strigolactones) and signalling molecules that attract beneficial fungi. In animals, carotenoids help to maintain health and promote reproduction. In humans, there is a daily dietary requirement for ingestion of carotenoids that are used to make vitamin A, which prevents eye diseases and some cancers.
Carotenoids are natural pigments synthesized by plants, algae, photosynthetic bacteria and aphids. In plants, carotenoids take role in photosynthesis, photoprotection and production of some phytohormones (Abscisic acid and Strigolactones) and signalling molecules that attract beneficial fungi. In animals, carotenoids help to maintain health and promote reproduction. In humans, there is a daily dietary requirement for ingestion of carotenoids that are used to make vitamin A, which prevents eye diseases and some cancers.
Through the use of genetic engineering, new biofortification strategies have been developed to enhance the access to these health promoting compounds. One of the best known examples is the enrichment of carotenoids in white rice to make golden rice. However, biofortification strategies can be unpredictable due to metabolic feedback regulation that maintains metabolite composition insuring cellular homeostasis. Therefore, a deeper understanding and knowledge of feedback regulatory mechanisms is required to advance genomic engineering approaches for successful biofortication.
In eukaryotic systems, gene transcription occurs when the information in specific segment of DNA copies itself into a messenger RNA (mRNA) which has 5' and 3' untranslated regions (5' & 3' UTR) either side of the protein coding region. The 5'UTR can affect gene transcription and protein translation depending upon developmental and environmental factors. We hypothesise that the 5'UTR is an important regulatory switch sensing metabolite levels and controlling the regulation of carotenogenesis in plants. Similar mechanisms can operate in bacteria to fine-tune metabolic levels in response to environmental stress.
When plant seeds germinate underground in the dark they soon become exposed to light triggering major changes in energy organelle biogenesis and secondary metabolic processes. This transition from dark to light provides an ideal model to investigate the effects of metabolite accumulation on feedback regulatory mechanisms in plants.
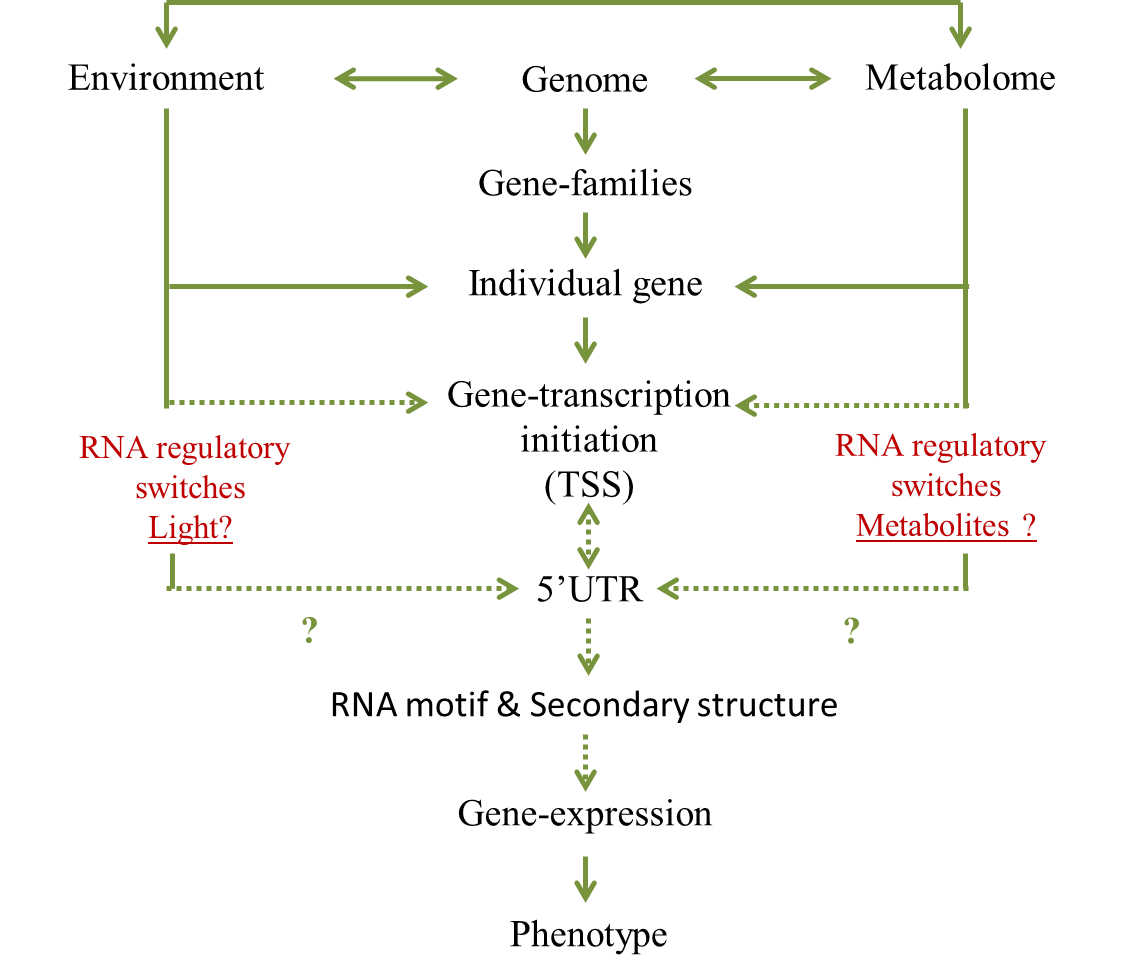
My research interests include understanding the genome-wide regulation of transcription start sites (TSS) and role of 5'UTR in the metabolic feedback regulation in carotenoid biosynthesis pathway (see Figures below). An ultimate goal of my research is to discover novel RNA regulatory switches in the 5'UTR of mRNA that control gene expression in response to environmental and developmental cues.
- Personal website: https://yagizalagoz.wordpress.com/ (opens in a new window)
- ORCID ID: http://orcid.org/0000-0001-5081-6874 (opens in a new window)
Awards
- 2018 Research Science Award (3rd Place) in COMBIO2018, Australian Society for Plant Scientists (ASPS), ICC Sydney (24th-26th Sept. 2018, 250AUD)
- 2018 Poster Award, Gordon Research Conference on Carotenoids, Maine, USA (16th-22nd June 2018, 500USD)
- 2018 Travel Grant for COMBIO2018, Australian Society for Plant Scientists (ASPS) (165AUD)
- 2017 Registration Support for Gene Editing of Crops Workshop, CSIRO, Australian Society of Plant Scientists (ASPS) (150AUD)
- 2016-2019 Hawkesbury Institute for the Environment-Postgraduate Research Award (HIE-PRA) with Tuition Fee Waiver Scholarship and OSHC, Western Sydney University (UWS) (Jan 11, 2016, AUD25849 per annum stipend)
- 2013-2015 Research Project Scholarship, The Scientific and Technological Research Council of Turkey (TUBITAK), International Olive Genome Consortium (IOGC) (TRY18000 per annum stipend)
- 2012 Erasmus Summer Internship Grant, European Commission (€1620).
- 2012 Travel Grant for Virus-Induced Gene Silencing-Training School (VIGS-TS), UK, Rothamsted Research, European Cooperation in Science and Technology (EU-COST) (€1000).
- 2011 Poster Award (2nd Place), Biotech2011 National Biotechnology Student Conference, Istanbul, Turkey.
- 2008-2013 Undergraduate Full Tuition Fee Waiver Scholarship, Council of Higher Education (CoHE) of Republic of Turkey (YÖK)
Publications
Book Chapter
Alagoz Y., Dhami N., Mitchell C., Cazzonelli CI. (2020) cis/trans Carotenoid extraction, purification, detection, quantification, and profiling in plant tissues. In: Concepción MR, Welsch R (eds) Plant and food carotenoids: methods and protocols. Springer, New York. doi:10.1007/978-1-4939-9952-11
Journals
Chavan SG, He X, Maier C, Alagoz Y, Anwar S, Chen Z-H, Ghannoum O, Cazzonelli CI, Tissue DT, (2023) 'An energy-saving glasshouse film reduces seasonal, and cultivar dependent Capsicum yield due to light limited photosynthesis', Annals of Agricultural Sciences, vol.68, no.1, pp 21 - 35
Anwar S, Brenya E, Alagoz Y, Cazzonelli CI, (2021) 'Epigenetic control of carotenogenesis during plant development', Critical Reviews in Plant Sciences, vol.40, no.1, pp 23-48
Cazzonelli CI, Hou X, Alagoz Y, Rivers J, Dhami N, Lee J, Marri S, Pogson BJ, (2020) 'A cis-carotene derived apocarotenoid regulates etioplast and chloroplast development', ELife, vol.9, Article no.e45310
Chavan SG, Maier C, Alagoz Y, Filipe JC, Warren CR, Lin H, Jia BH, Loik ME, Cazzonelli CI, Chen ZHH, Ghannoum O, Tissue DT, (2020) 'Light-limited photosynthesis under energy-saving film decreases eggplant yield', Food and Energy Security, Article no.e245
Alagoz Y, Nayak P, Dhami N, Cazzonelli CI, (2018) 'cis-carotene biosynthesis, evolution and regulation in plants: The emergence of novel signaling metabolites', Archives of Biochemistry and Biophysics, vol.654, pp 172-184
Alagoz Y, Gurkok T, Parmaksiz I, Unver T, (2016) 'Identification and Sequence Analysis of Alkaloid Biosynthesis Genes in Papaver Section Oxytona', Turkish Journal of Biology, vol.40, pp 174-183
Alagoz Y, Gurkok T, Zhang B, Unver T, (2016) 'Manipulating the Biosynthesis of Bioactive Compound Alkaloids for Next-Generation Metabolic Engineering in Opium Poppy Using CRISPR-Cas9 Genome Editing Technology', Scientific Reports, vol.6, Article no.30910
Research supervisors
Dr Christopher Cazzonelli, Professor David Tissue, Dr Alexie Papanicolaou